 |
SMILE
v2.5
Schwarzschild Modelling Interactive expLoratory Environment
|
 |
SMILE
v2.5
Schwarzschild Modelling Interactive expLoratory Environment
|
frozen N-body potential calculated by tree-code algorithm (based on hackcode1.c) More...
#include <potential.h>
Classes | |
| struct | cell |
| CELL: structure used to represent internal nodes of tree. More... | |
| struct | node |
| NODE: data common to BODY and CELL structures. More... | |
Public Member Functions | |
| CPotentialNB () | |
| default (empty) initialization | |
| template<typename NumT > | |
| CPotentialNB (double _eps, double _tol, const CPointMassSet< NumT > &points) | |
| Initialize from a list of particles;. More... | |
| virtual CPotential * | clone () const |
| Return a pointer to a copy of this instance of potential. More... | |
| virtual POTENTIALTYPE | PotentialType () const |
| enumerable potential type | |
| virtual const char * | PotentialName () const |
| string representation of potential type | |
| virtual SYMMETRYTYPE | symmetry () const |
| returns symmetry type of this potential | |
| virtual double | Phi (double X, double Y, double Z, double t=0) const |
| Return potential at a given spatial point (possibly a time-varying one). More... | |
| virtual double | Rho (double X, double Y, double Z, double t=0) const |
| returns density at given coordinates, this should obviously be overriden in derivative classes | |
| virtual void | Force (const double xyz[N_DIM], const double t, double *force, double *forceDeriv=NULL) const |
| Compute forces and, optionally, force derivatives at a given point. More... | |
| virtual double | Mass (const double r) const |
| reimplement the mass estimate by counting bodies inside given radius | |
| size_t | bodyNum () const |
| return number of particles | |
| template<typename NumT > | |
| CPosPoint< NumT > | bodyPos (size_t index) const |
| return the position of a given particle | |
| double | bodyMass (size_t index) const |
| return the mass of a given particle | |
 Public Member Functions inherited from smile::CDensity Public Member Functions inherited from smile::CDensity | |
| virtual double | totalMass () const |
| returns estimated M(r=infinity) or -1 if mass is infinite | |
| double | getRadiusByMass (const double m) const |
| solves for Mass(r)=m | |
| void | getRadiiByMass (const vectord &masses, vectord *radii) const |
| solves for Mass(r)=m for an array of sorted values of m (more efficient than doing it one-by-one) | |
| bool | checkMassMonotonic () const |
| safety measure: check (roughly) that mass is increasing with radius | |
| bool | checkDensityNonzero () const |
| another safety measure: check that density doesn't drop to zero along any of three axes (important to assess spherical-harmonic approximation quality) | |
| virtual double | getGamma () const |
| returns inner density slope estimate (only used in BSE potential expansion for the automatic selection of shape parameter Alpha) | |
Static Public Member Functions | |
| static const char * | myName () |
Private Types | |
| enum | NODETYPE { BODY, CELL } |
| tree contains nodes of two kinds: individual particles and child cells More... | |
| enum | WALKOP { CALCPHI, CALCACC } |
| operation to perform during the tree-walk More... | |
| typedef double | vec3 [N_DIM] |
| typedef double | matrix3 [N_DIM][N_DIM] |
| typedef node * | nodeptr |
| typedef node | body |
| BODY: data structure used to represent particles. | |
| typedef node * | bodyptr |
| typedef cell * | cellptr |
Private Member Functions | |
| void | maketree () |
| initialize tree for the constructor | |
| bool | expandbox (const size_t b) |
| expand bounding box (aka root cell) if particle does not fit into existing one | |
| bool | appendtree (const size_t b) |
| append particle to the tree | |
| bool | intcoord (int xp[N_DIM], const vec3 rp) const |
| calculate integerized coords | |
| int | subindex (const int x[N_DIM], const int l) const |
| determine subcell that a given position belongs to (l is cell level) | |
| void | centerofmass (const nodeptr q, const vec3 corner, const double cellsize, int l) |
| recursive calculation of cell center-of-mass | |
| void | assigneps (const nodeptr q, const double epsparent, const double cellsize) |
| recursive initialization of individual softening radii of cells More... | |
| void | propagateeps (const nodeptr q) |
| recursively assign epsilon to tree cells from underlying nodes | |
| void | walktree (const nodeptr p, const double dsq, const vec3 pos0, const WALKOP operation, double data[]) const |
| recursive computation of potential or forces from tree node/cell p at position pos0 | |
| void | walktreedens (const nodeptr p, const double dsq, const double searchradsq, const vec3 pos0, const int xp[N_DIM], const int lev, vectorpd *data) const |
| recursive search of particles in node/cell p which are closer to pos0 than sqrt{dsq). More... | |
Private Attributes | |
| const double | eps |
| potential softening parameter; negative value means position-dependent softening proportional to local density^{-1/3} | |
| const double | tol |
| accuracy parameter (opening angle): 0 means exact N^2 force calculation (don't ever try!) | |
| std::vector< body > | bodytab |
| list of all particles | |
| std::vector< cell > | celltab |
| list of all tree cells | |
| nodeptr | troot |
| pointer to the root cell of the tree | |
| vec3 | rmin |
| lower-left corner of coordinate box | |
| double | rsize |
| side-length of integerized coordinate box | |
Static Private Attributes | |
| static const int | IMAX =(1 << (8 * sizeof(int) - 2)) |
| max integer coordinate | |
| static const unsigned int | NSUB =(1 << N_DIM) |
| subcells per cell (2^3) | |
Additional Inherited Members | |
 Public Types inherited from smile::CDensity Public Types inherited from smile::CDensity | |
| enum | POTENTIALTYPE { PT_UNKNOWN, PT_DIRECT, PT_COMPOSITE, PT_COEFS, PT_NB, PT_BSE, PT_BSECOMPACT, PT_SPLINE, PT_CYLSPLINE, PT_LOG, PT_HARMONIC, PT_SCALEFREE, PT_SCALEFREESH, PT_SPHERICAL, PT_DEHNEN, PT_MIYAMOTONAGAI, PT_FERRERS, PT_PLUMMER, PT_ISOCHRONE, PT_PERFECTELLIPSOID, PT_NFW, PT_SERSIC, PT_EXPDISK, PT_ELLIPSOIDAL, PT_MGE } |
| list of all existing types of density or density/potential models, each of them implemented in its own class More... | |
| enum | SYMMETRYTYPE { ST_NONE = 0, ST_REFLECTION = 1, ST_PLANESYM = 2, ST_ZROTSYM = 4, ST_SPHSYM = 8, ST_TRIAXIAL = ST_REFLECTION | ST_PLANESYM, ST_AXISYMMETRIC = ST_TRIAXIAL | ST_ZROTSYM, ST_SPHERICAL = ST_AXISYMMETRIC | ST_SPHSYM, ST_DEFAULT = ST_TRIAXIAL } |
| Type of symmetry. More... | |
frozen N-body potential calculated by tree-code algorithm (based on hackcode1.c)
|
private |
|
private |
| smile::CPotentialNB::CPotentialNB | ( | double | _eps, |
| double | _tol, | ||
| const CPointMassSet< NumT > & | points | ||
| ) |
Initialize from a list of particles;.
| NumT | = double or float |
|
private |
recursive initialization of individual softening radii of cells
recursive assign of softening length: compute the number density of points in a given cell and average it with the parent cell, repeat process for the entire tree is scanned several times to perform back and forth averaging over parent/daughter nodes.
eps_i = |eps| * n^{-1/3}, where eps is the global softening parameter and n is the average number density in the vicinity of the node. epsparent is the softening length of the parent cell (zero on the first iteration).
|
inlinevirtual |
Return a pointer to a copy of this instance of potential.
A standard copy constructor or assignment is disabled because of different amount of data needed to be copied in different derived classes).
Implements smile::CPotential.
|
virtual |
Compute forces and, optionally, force derivatives at a given point.
| [in] | xyz | - coordinates of the point to compute forces (array of 3 numbers) |
| [in] | t | - time to compute forces (matters only if potential is time-dependent) |
| [out] | force | - computed values of -d Phi/d x_i; output array must exist and contain N_DIM values. |
| [out] | forceDeriv | - if not NULL, then also compute the second derivatives of potential (which is a symmetric matrix of size N_DIM^2, thus the array must be of size N_DIM*(N_DIM+1)/2 ): first 3 values contain 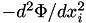 , second three contain mixed derivatives , second three contain mixed derivatives 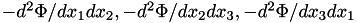 . . |
Implements smile::CPotential.
|
virtual |
Return potential at a given spatial point (possibly a time-varying one).
Implements smile::CPotential.
|
private |
recursive search of particles in node/cell p which are closer to pos0 than sqrt{dsq).
the list of such particles is stored in data and then used to compute density via kernel estimate
 1.8.8
1.8.8