 |
SMILE
v2.5
Schwarzschild Modelling Interactive expLoratory Environment
|
 |
SMILE
v2.5
Schwarzschild Modelling Interactive expLoratory Environment
|
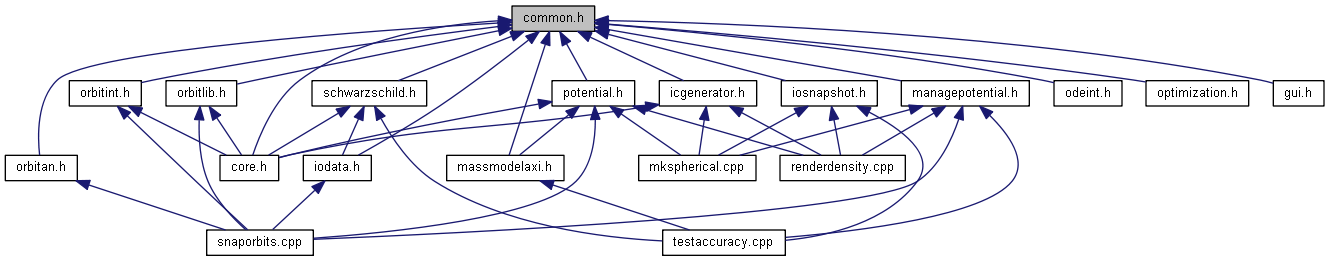
provides forward class declarations, several abstract data classes, a couple of global functions, and global constants (header file only). More...
#include <utility>#include <vector>#include <string>
Classes | |
| struct | smile::CPosPoint< NumT > |
| Data-only class containing 3 coordinates. More... | |
| struct | smile::CPosVelPoint< NumT > |
| Data-only class containing 3 coordinates and 3 velocities. More... | |
| struct | smile::CPosVelPoint< NumT > |
| Data-only class containing 3 coordinates and 3 velocities. More... | |
| struct | smile::CPosPoint< NumT > |
| Data-only class containing 3 coordinates. More... | |
| struct | smile::CPointMassSet< NumT > |
| An array of particles with positions, velocities and masses. More... | |
| struct | smile::CMatrix< NumT > |
| interface for matrix storage. More... | |
| struct | smile::CMatrixDense< NumT > |
| matrix interface class that actually holds the entire matrix and returns its elements More... | |
| struct | smile::CMatrixDiagonal< NumT > |
| matrix interface class that provides a diagonal matrix More... | |
| class | smile::CBasicQuadraticOptimizationSolver< NumT > |
| The interface class for various third-party quadratic optimization solvers. More... | |
| class | smile::CBasicOrbitFilteringFnc |
| Parent class for orbit filtering functions which evaluate whether an orbit should be used (in some calculation). More... | |
| class | smile::CBasicOrbitRuntimeFnc |
| defines a routine which may be called after each integration timestep to perform user-specific data collection. More... | |
| class | smile::CBasicOrbitRuntimeFncCreator |
| defines a basic class that creates some sort of runtime function for any orbit in the orbit library. More... | |
| class | smile::CBasicInformation |
| parent class for data container objects used to store various data and exchange it between objects. More... | |
| class | smile::COdeSystem |
| basic class of ODE system used in the orbit integrator. More... | |
| class | smile::CBasicOdeIntegrator |
| basic class for numerical integrators of ODE system More... | |
Namespaces | |
| smile | |
| common namespace for all core SMILE classes, functions and variables | |
Typedefs | |
------- define several convenience types ------- | |
| typedef float | smile::sdNumType |
| how to store orbit data for Schwarzschild modelling (float or double) | |
| typedef float | smile::smNumType |
| how to handle Schwarzschild modelling data internally (float or double) | |
| typedef std::vector< double > | smile::vectord |
| vector of double values | |
| typedef std::vector< float > | smile::vectorf |
| vector of float values | |
| typedef std::pair< double, double > | smile::paird |
| a pair of doubles | |
| typedef std::vector< paird > | smile::vectorpd |
| vector of pairs of doubles | |
| typedef std::pair< float, float > | smile::pairf |
| a pair of floats | |
| typedef std::vector< pairf > | smile::vectorpf |
| vector of pairs of floats | |
| typedef vectord | smile::OdeStateType |
| container for the array of variables used in ODE integration | |
Variables | |
----------- Parameters for potential and force evaluation ------------- | |
| const unsigned int | smile::N_DIM =3 |
| Dimensions of the real world. DO NOT CHANGE (unless you are God)! | |
| const size_t | smile::MAX_NCOEFS_RADIAL =101 |
| max number of basis-function expansion members (radial and angular). | |
| const size_t | smile::MAX_NCOEFS_ANGULAR =51 |
| const double | smile::EPSREL_DENSITY_INT =1e-4 |
| relative accuracy of density computation | |
| const double | smile::EPSREL_POTENTIAL_INT =1e-6 |
| relative accuracy of potential computation (integration tolerance parameter) | |
| const double | smile::EPSABS_POTENTIAL_INT =1e-15 |
| absolute error in potential computation (supercedes the relative error in case of very small coefficients) | |
| const double | smile::EPSREL_MASS_INT =1e-2 |
| relative accuracy of cell mass computation in Schwarzschild modelling | |
| const double | smile::MIN_RADIUS_CUTOFF =1e-6 |
| inner and outer cutoff radii appearing in calculations of M(r), r(E) and related | |
| const double | smile::MAX_RADIUS_CUTOFF =1e+6 |
| const double | smile::SPLINE_MIN_RADIUS =1e-10 |
| auxiliary softening parameter for Spline potential to avoid singularities in some odd cases | |
------------- Orbit integration parameters ---------- | |
| #define | DOP853_DENSE_OUTPUT_FAST |
| for DOP853 integrator, setting this flag eliminates three calls to force function per timestep at the expense of getting a slightly lower (still acceptable) accuracy in the interpolated solution (6th instead of 7th order) | |
| #define | TREECODE_QUADRUPOLE 1 |
| whether to use quadrupole moments in tree expansion of Nbody potential (no reason to switch it off) | |
| #define | TREECODE_EPS_COMPACT_KERNEL 1 |
| if defined, use compact Epanechnikov kernel for potential/force softening, otherwise use Plummer softening (not recommended?) | |
| #define | TREECODE_EPS_FROM_DENSITY 0 |
| if defined, evaluate adaptive softening using true local density, otherwise use a more approximate but faster method based on cell volumes | |
| #define | TREECODE_DEHNEN_MAC 0 |
| if defined, use Dehnen's (2000) multipole acceptance criterion based on r_max - max distance between cell's c.o.m and its child nodes, otherwise use standard Barnes&Hut criterion based on cell size | |
| #define | TREECODE_DENSITY_SPLINE 0 |
| method to smooth density: standard SPH spline, or Epanechnikov kernel (both have finite support equal to distance to Kth neighbour) | |
| const size_t | smile::NUMSTEP_MAX =static_cast<size_t>(1e7) |
| maximal number of integration steps (in case something went wrong with the timestep, do not dead-lock) | |
| const double | smile::TREECODE_TIMESTEP_BH_FACTOR =0.2 |
| for Nbody treecode, another safety measure is to decrease timestep near BH by this factor (in the leap-frog integrator) | |
| const size_t | smile::TREECODE_DENSITY_KTH_NEIGHBOUR =32 |
| density estimate in treecode by using distance to k-th neighbour; if using spline kernel, put more (>~20), since this kernel is more concentrated | |
| const double | smile::LYAP_DEV_INIT =1e-10 |
| initial value of orbit separation (if variational equation is not used and two nearby trajectories are followed) | |
| const double | smile::LYAP_DEV_MAX =1e-6 |
| if distance between nearby trajectories reaches the following threshold, renormalize it back to LYAP_DEV_INIT | |
| const double | smile::LYAP_VAREQ_DEV_MAX =1e5 |
| if variational equation is used, perform renormalization of deviation vector if greater than threshold (only for numerical convenience, since the var.equation is anyway linear) | |
------------- Orbit spectral analysis and classification options ---------- | |
| #define | FREQ_DIFF_METHOD 2 |
| method to estimate frequency diffusion rate. More... | |
| const size_t | smile::MAX_FFT_PRIME =41 |
| orbit spectrum is calculated by fast fourier transform from GSL. More... | |
| const double | smile::FREQ_PRECISE_THRESHOLD =5.0 |
| Use Hunter's method for precise determination of frequency of a given line if the spectrum neat its maximum is reasonably similar to that of a single line. More... | |
| const size_t | smile::MAX_SPECTRAL_LINES =10 |
| maximal number of spectral lines in each coordinate to search for | |
| const double | smile::MIN_SPECTRAL_LINE_AMPLITUDE =1e-2 |
| do not use lines which amplitude is X times lower than amplitude of the leading line | |
| const double | smile::FREQ_ACCURACY =0.1 |
| accuracy of frequency comparison (to determine resonances), in units of Nyquist frequency | |
| const double | smile::FREQ_DIFF_REG_MIN =1e-6 |
| minimum value of freq.diff threshold separating regular and chaotic orbits (actual threshold depends on integration interval) | |
| const int | smile::FREQ_RES_2 =10 |
| maximum order of m:n frequency resonance (or linear dependence of three frequencies) | |
| const int | smile::FREQ_RES_3 =10 |
| maximum order of l*a+m*b+n*c frequency commensurability (thin orbit) | |
| const double | smile::OC_PYRAMID_AVGZ =0.1 |
| threshold value of |<z>| / rmax to classify orbit as pyramid or saucer | |
| const size_t | smile::MAX_POINCARE_POINTS =10000 |
| max number of points in Poincare section | |
| const double | smile::PERICENTER_FIT_FRACTION =0.1 |
| fraction of closest pericenter passages which are used in fitting | |
| const int | smile::PERICENTER_FIT_MIN_POINTS =10 |
| minimum number of points used in pericenter fitting | |
| const int | smile::PERICENTER_FIT_MAX_POINTS =50 |
| defines max points in pericenter fitting | |
------------- Orbit library / Schwarzschild modelling options --------------- | |
sample initial conditions out to a radius containing this fraction of mass | |
| #define | SCHWARZSCHILD_START_SPACE 1 |
| number of significant digits for initial conditions (rounded to this accuracy) More... | |
| #define | NUM_SUBDIV_ANGULAR 5 |
| the mass in each cell of the classic Schwarzschild grid is computed by dividing it into a number of smaller subcells in two angular directions; this number is given below | |
| const double | smile::MAX_LAGRANGIAN_RADIUS =0.999 |
| const size_t | smile::NUM_RADIAL_POINTS_SPHERICAL_MODEL =100 |
| number of radial points used in createMassModel | |
| const double | smile::SCHW_MAX_CELLMASS_REL_DIFFERENCE =1e-2 |
| max relative difference in cell mass to consider the cell as feasible: abs(1-Mcell/Mcellrequired) | |
| const double | smile::SCHW_MIN_ORBIT_WEIGHT_USED =1e-4 |
| min orbit weight to use the orbit after the optimization problem solved (divide by number of orbits!) | |
| const double | smile::NBEXPORT_XV_FACTOR =10.0 |
| Nbody export: relative importance of nullifying total angular momentum at the expense of pericenter velocity * pericenter offset. | |
-------------- Options for GUI and high-level operations ------------ | |
| #define | SMILEVERSION "2.5" |
| SMILE version identifier. | |
| #define | USE3DFACETS |
| whether to perform triangulation and display 'envelope' of an orbit (useless without Qwt3D; requires external program QDelaunay from QHull package) | |
| #define | DEFAULT_SETTINGS_FILE_NAME "smile.ini" |
| auto-load settings from file | |
| const size_t | smile::NUM_POINTS_PLOT_EQUIPOTENTIAL =100 |
| number of points in curve representing equipotential surface (actually *4) | |
| const size_t | smile::NUM_POINTS_PLOT_POINCARE =100 |
| number of points in outer bounding curve in Poincare section (actually *4) | |
| const double | smile::SCHW_MIN_ORBIT_WEIGHT_DISPLAYED =1e-4 |
| min. weight of orbit to be displayed in GUI | |
| const double | smile::PLOT_FREQ_MAX_F =5.0 |
| width of frequency spectrum displayed (in units of orbital frequency) | |
-------- Various convenience definitions -------- | |
|
typedef std::vector < CBasicOrbitRuntimeFnc * > | smile::vectorRuntimeFncs |
| convenience definition of vectors of timestep functions and function creators | |
|
typedef std::vector< const CBasicOrbitRuntimeFncCreator * > | smile::vectorRuntimeFncCreators |
|
typedef std::vector< const CBasicInformation * > | smile::vectorInformation |
|
typedef std::vector < CPosVelPoint< double > > | smile::CPosVelDataDouble |
| convenience definitions of templated vectors containing particle sets with/without masses | |
|
typedef std::vector < CPosVelPoint< float > > | smile::CPosVelDataFloat |
| double | smile::pow_2 (double x) |
| convenience shorthand for squaring a number | |
| float | smile::pow_2 (float x) |
| size_t | smile::pow_2 (size_t x) |
| template<typename T1 , typename T2 > | |
| bool | smile::comparepair (const std::pair< T1, T2 > &val1, const std::pair< T1, T2 > &val2) |
| convenience function for sorting arrays of std::pair<float/double, something> | |
provides forward class declarations, several abstract data classes, a couple of global functions, and global constants (header file only).
| #define FREQ_DIFF_METHOD 2 |
method to estimate frequency diffusion rate.
0 - difference in the largest of three frequencies (Valluri&Merritt 98), 1 - average of differences in all three freq, 2 - average over those coords for which |f1-f2|/(f1+f2)<0.5 (more robust to spurious errors in lf determination)
| #define SCHWARZSCHILD_START_SPACE 1 |
number of significant digits for initial conditions (rounded to this accuracy)
if using Schwarzschild stationary start space (3d), put points on a square grid in 3 sectors adjacent to X, Y and Z axes (same as in Schwarzschild 1993). otherwise, put points along lines of constant Z (=constant theta), number of points being proportional to sin(theta) (same as in Terzic 2002).
 1.8.8
1.8.8